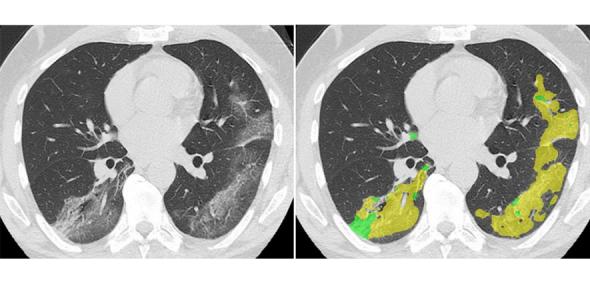
An open-source artificial intelligence (AI) tool, combining chest imaging data with laboratory and clinical data, is being developed by Cambridge researchers to support the rapid diagnosis and triaging of patients with COVID-19 in the UK.
The team, led by Professors Carola-Bibiane Schönlieb and Evis Sala, brings together expertise in AI for imaging with expertise in radiology and clinical applications from Addenbrooke’s and Papworth Hospitals, as well as collaborators from the UK, China, Austria and Italy, to develop a prediction model that can rapidly and reliably diagnose and suggest a prognosis to doctors.
Reverse-transcription polymerase chain reaction (RT-PCR) tests are currently the most common tool used to diagnose COVID-19, but they are only up to 70% sensitive, meaning there are up to 30% false negatives.
While chest X-rays and CT scans provide valuable diagnostic and monitoring information that can complement laboratory and clinical data, it is a complex task typically done by radiologists, whose expertise is often in high demand. Fast and accurate diagnosis of patients in order to limit disease spread, together with the rapid determination of whether a patient is likely to recover, require intensive care unit (ICU) admission, or intensive ventilation, is key to allocating resources and to improving patient outcomes.
“AI offers huge potential to support agile clinical decision making, ensuring patients receive the most appropriate support and leading to better patient outcomes,” said Sala, who is based at the Department of Radiology.
Recent studies have suggested that using AI could have a meaningful impact on the management of patients with COVID-19. AI tools such as deep learning can offer automated image analysis and integration with clinical data to help clinicians make more informed decisions for treatment.
However, good quality data and computing power are required to train and optimise predictive AI models and data availability is a major bottleneck when developing new systems. Coupled with this, the lack of standardisation of datasets makes it challenging to reuse existing AI tools in a different country than the one that it was trained for. Most current AI tools have been developed on small, locally collected datasets. Data that is being collected in hospitals all over the world varies in what is being collected and how the data is processed. Therefore, an effort for developing a widely applicable tool for COVID-19 hospital support must be open source so it can be adapted to different environments; be based on a serious data sharing and data curation, data cleaning and standardisation effort; and be developed with mathematical, statistical and engineering expertise to develop robust and translatable tools.
To address these challenges, the team from the Cambridge Centre for Mathematical Imaging in Healthcare (CMIH) is developing a flexible, open-source AI tool that could be used by hospitals worldwide. Drawing on their history of global research collaboration and expertise in data governance, the team is gathering datasets from Austria, China, Italy and the UK for their work. Data scientists and clinicians are working in close collaboration, following standard protocols to identify bias during development. “Rigorous mathematical models play a key role in mitigating bias and improving the efficacy of the prediction model as they follow universal rules with mathematical guarantees,” said Schönlieb.
Using deep learning approaches along with mathematical and statistical analysis methods, the new tool will be accompanied by a comprehensive algorithmic strategy that will allow fine-tuning for datasets with different characteristics and implementation in different countries. The team are hoping to launch the AIX-COV-NET tool within the next 12 to 18 months. The project has recently received funding from the EU-funded Innovative Medicines Initiative and Intel.
“Our team’s strength is the close dialogue we have between clinicians and data scientists, and the passion we all bring for advancing AI solutions for COVID-19,” said Schönlieb, from Cambridge's Department of Applied Mathematics and Theoretical Physics.
“AI offers huge potential to support agile clinical decision making, ensuring patients receive the most appropriate support and leading to better patient outcomes,” said Sala.
The core project team is comprised of data scientists and clinicians from across Cambridge and is led by Professor Carola-Bibiane Schönlieb, Director of the Centre for Mathematical Imaging in Healthcare, and Professor Evis Sala, Professor of Oncological Imaging, University of Cambridge & Honorary Consultant Radiologist, Addenbrooke’s Hospital, Cambridge University Hospitals NHS Foundation Trust. The team is supported by an AI and image analysis team, drawn from subject experts across Cambridge and around the world, a clinical team comprised of colleagues from hospitals in Cambridge, London and Vienna, and a support team based in the Faculty of Mathematics. Partner institutions include hospitals in Wuhan, China; Milan, Italy; and Madrid, Spain, and universities in Manchester, Vienna and London.
The University of Cambridge has an impressive record of achievement in multidisciplinary research and innovation. The CMIH is a collaboration between mathematics, engineering, physics and biomedical scientists and clinicians and is one of five centres to receive investment from the Engineering and Physical Sciences Research Council (EPSRC). A key aim of this partnership is the delivery of high quality, multidisciplinary research that will help overcome some of the major challenges facing the NHS.
How you can support Cambridge's COVID-19 research effort
Donate to support COVID-19 research at Cambridge
AI assisted COVID-19 diagnostic and prognostic tool could improve resource allocation and patient outcomes.

The text in this work is licensed under a Creative Commons Attribution 4.0 International License. Images, including our videos, are Copyright ©University of Cambridge and licensors/contributors as identified. All rights reserved. We make our image and video content available in a number of ways – as here, on our main website under its Terms and conditions, and on a range of channels including social media that permit your use and sharing of our content under their respective Terms.















