Dr Phil Jones
Position: Senior Group Leader
Personal home page:
http://www.sanger.ac.uk/people/directory/jones-phil-h
PubMed journal articles - click here
Dr Phil Jones is pleased to consider applications from prospective PhD students.
Our group studies how normal cell behaviour is altered by mutation in the early stages of cancer evolution. We focus on squamous tissues, the skin epidermis and the lining of the oesophagus, using transgenic models, novel sequencing approaches in human tissues, live imaging and single cell analysis to uncover key steps in cancer development, with aim of developing rational interventions to decrease cancer risk.
Cell dynamics in normal and mutant stem cells
Our group pioneered the use of large scale genetic lineage tracing to quantify cell behaviour in vivo. This approach reveals that the normal cell turnover of the epidermis and oesophageal epithelium is maintained by a single population of progenitor cells. Progenitor division generates two progenitor daughters, two non-dividing differentiated cells or one progenitor and one differentiating cell. The outcome of a given division is unpredictable, but in homeostasis the probabilities of producing two progenitor and two differentiating daughters are the same, so that on average, equal numbers of progenitors and differentiating cells are produced across whole population of progenitors. Following injury, nearby progenitors switch from 'maintenance mode' to produce an excess of progenitor daughters, until the tissue is repaired, when they revert to maintenance mode once more. This behavioural plasticity allows tissue repair without the need for 'reserve' stem cells.
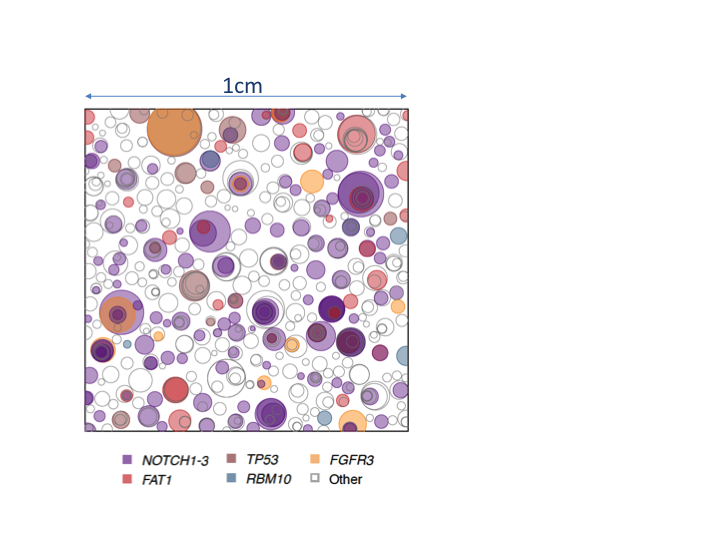
The ability of progenitors to produce excess progenitor daughters for wound repair represents a potential vulnerability. Mutations may bias progenitor cell fate towards proliferation. We have shown that both p53 and Notch inhibiting mutations imbalance cell fate in this way, generating large mutant clones. We have developed a sequencing based approach to show similar in human tissues, finding around a third of cells in normal sun exposed facial skin in middle aged caucasians carry cancer driver gene mutations. Remarkably, normal tissues restrain the expansion of mutant clones, so very few of them progress to form tumours. One focus of our current research is to understand the mechanism of this restraint. We are also investigating mutant clones in other human tissues.
The molecular basis of progenitor cell fate decisions
We are using live cell imaging, single cell analysis and gene editing to resolve the molecular deteminants of human epidermal progenitor cell division outcomes and the mode of cell division, guided by the mutations which drive clonal expansion in normal human skin.
Symplectic Elements feed provided by Research Information, University of Cambridge
Human keratinocytes have two interconvertible modes of proliferation. Roshan A, Murai K, Fowler J, Simons BD, Nikolaidou-Neokosmidou V and Jones PH
Nature cell biology 2016; 18;145-156. Tumor evolution. High burden and pervasive positive selection of somatic mutations in normal human skin. Martincorena I, Roshan A, Gerstung M, Ellis P, Van Loo P et al. Science 2015;348;880-6. Differentiation imbalance in single oesophageal progenitor cells causes clonal immortalization and field change. Alcolea MP, Greulich P, Wabik A, Frede J, Simons BD and Jones PH. Nature cell biology 2014;16;615-22.